Modeling Mortality Trends with Functional Forecasting
Source:vignettes/mortality-analysis.Rmd
mortality-analysis.RmdIntroduction
Mortality patterns represent one of the most fundamental demographic
processes, yet traditional statistical approaches often miss the
complex, evolving functional relationships between age and death rates
over time. This vignette demonstrates how functional forecasting using
the ffc package captures these dynamic relationships that
change shape—not just magnitude—over time.
What makes mortality data ideal for functional forecasting?
- Complex age patterns: Mortality exhibits characteristic J-shaped curves across age groups
- Temporal evolution: These functional relationships shift
systematically over time
- Smooth transitions: Changes occur gradually, making them suitable for GAM-based smoothing
- Forecasting relevance: Understanding future mortality trends has critical policy implications
Key concepts we’ll explore
- Time-varying coefficients: How functional relationships evolve over time
- Hierarchical smoothing: Modeling shared trends with group-specific
deviations
- Functional forecasting: Predicting entire curves into the future
- Model diagnostics: Validating functional time series models
Data exploration
We’ll use Queensland mortality data spanning 1980-2020, containing death counts by age, sex, and year with corresponding population denominators.
data("qld_mortality")
head(qld_mortality, 15)
#> # A tsibble: 15 x 5 [1Y]
#> # Key: age, sex [1]
#> year age sex deaths population
#> <int> <int> <fct> <dbl> <dbl>
#> 1 1980 0 female 190 17700.
#> 2 1981 0 female 175 18785.
#> 3 1982 0 female 190 19698.
#> 4 1983 0 female 165 19908.
#> 5 1984 0 female 148 19573.
#> 6 1985 0 female 164 19458.
#> 7 1986 0 female 147 19405.
#> 8 1987 0 female 159 19421.
#> 9 1988 0 female 153 19841.
#> 10 1989 0 female 184 20942.
#> 11 1990 0 female 155 21841.
#> 12 1991 0 female 143 22187.
#> 13 1992 0 female 154 22314.
#> 14 1993 0 female 149 22581.
#> 15 1994 0 female 109 22859.The dataset structure is ideal for functional analysis: we have a functional predictor (age) whose relationship with the outcome (mortality) changes over time (year), with hierarchical structure (sex).
Visualizing mortality patterns
ggplot(
data = qld_mortality,
aes(
x = age,
y = deaths / population,
group = year,
colour = year
)
) +
geom_line(alpha = 0.7) +
facet_wrap(~sex) +
scale_colour_viridis_c(name = "Year") +
labs(
x = "Age",
y = "Mortality rate",
title = "Queensland mortality patterns over time"
) +
scale_y_log10()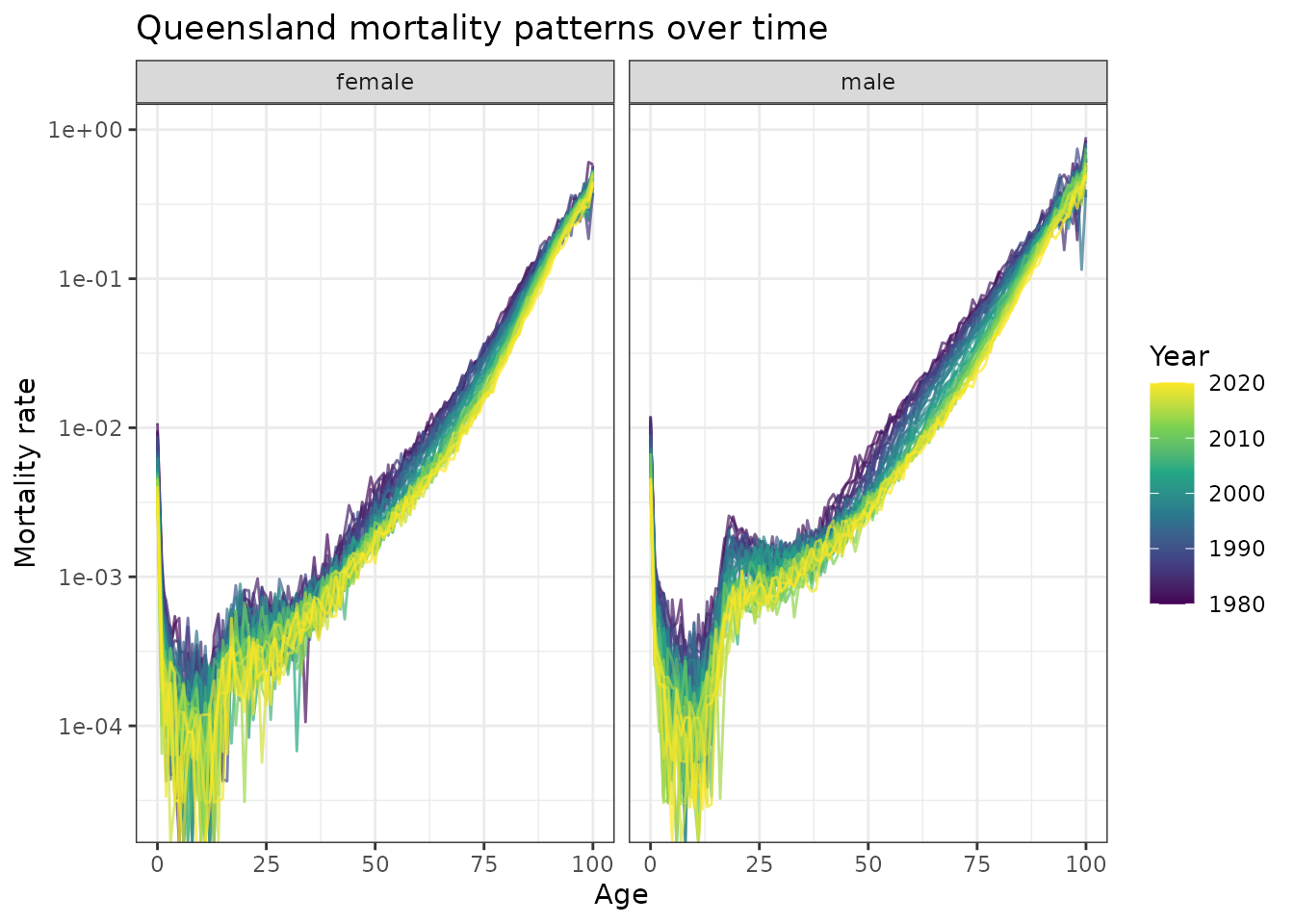
Observed mortality rates by age in Queensland, 1980-2020. The characteristic J-shaped curves show systematic downward shifts over time, indicating mortality improvements across all age groups.
Key observations:
- J-shaped curves: High infant mortality, low childhood mortality, exponential increase with age
- Temporal shifts: Consistent downward movement of entire curves over
time
- Sex differences: Males consistently higher mortality, similar temporal patterns
- Functional evolution: The shape itself evolves, not just vertical shifts
- Overall decline: Mortality rates have declined substantially across all ages, reflecting major improvements in healthcare, lifestyle, and living conditions
Functional modeling approach
Why traditional models fall short
Standard approaches might model this as:
# Traditional approach - misses functional evolution
glm(deaths ~ age + year + sex, family = poisson(), offset = log(population))This assumes linear age effects and additive time trends—clearly inadequate for evolving functional relationships.
The ffc solution: time-varying coefficients
The fts() function creates basis functions whose
coefficients evolve over time, capturing how the age-mortality
relationship changes:
mod <- ffc_gam(
deaths ~
offset(log(population)) +
sex +
age +
# Time-varying level: shared temporal trends
fts(
year,
mean_only = TRUE,
bs = "tp",
time_k = 35,
time_m = 1
) +
# Time-varying age effects: sex-specific deviations
fts(
age,
by = sex,
bs = "tp",
time_k = 15,
time_m = 1
),
time = "year",
data = qld_mortality,
family = poisson(),
# Efficient for large datasets
engine = "bam"
)Understanding the model structure
summary(mod)
#>
#> Family: poisson
#> Link function: log
#>
#> Formula:
#> deaths ~ sex + age + offset(log(population)) + s(year, by = fts_year1_mean,
#> bs = "ts", k = 35, m = 1, id = 1) + s(year, by = fts_bs_s_age_bysexfemale_1,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexfemale_2,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexfemale_3,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexfemale_4,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexfemale_5,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexfemale_6,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexfemale_7,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexfemale_8,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexfemale_9,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexmale_1,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexmale_2,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexmale_3,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexmale_4,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexmale_5,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexmale_6,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexmale_7,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexmale_8,
#> bs = "ts", k = 15, m = 1, id = 2) + s(year, by = fts_bs_s_age_bysexmale_9,
#> bs = "ts", k = 15, m = 1, id = 2)
#>
#> Parametric coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 3.371181 2.488125 1.355 0.175447
#> sexmale 0.574306 0.004986 115.189 < 2e-16 ***
#> age -0.180386 0.049764 -3.625 0.000289 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Approximate significance of smooth terms:
#> edf Ref.df Chi.sq p-value
#> s(year):fts_year1_mean 32.048 34 10329.34 <2e-16 ***
#> s(year):fts_bs_s_age_bysexfemale_1 14.470 15 11934.09 <2e-16 ***
#> s(year):fts_bs_s_age_bysexfemale_2 12.621 15 9711.83 <2e-16 ***
#> s(year):fts_bs_s_age_bysexfemale_3 14.477 15 68.77 <2e-16 ***
#> s(year):fts_bs_s_age_bysexfemale_4 13.307 15 7609.29 <2e-16 ***
#> s(year):fts_bs_s_age_bysexfemale_5 14.505 15 498.50 <2e-16 ***
#> s(year):fts_bs_s_age_bysexfemale_6 13.147 15 7787.91 <2e-16 ***
#> s(year):fts_bs_s_age_bysexfemale_7 14.265 15 712.25 <2e-16 ***
#> s(year):fts_bs_s_age_bysexfemale_8 9.615 15 11101.16 <2e-16 ***
#> s(year):fts_bs_s_age_bysexfemale_9 11.815 15 39.30 <2e-16 ***
#> s(year):fts_bs_s_age_bysexmale_1 14.347 15 7385.09 <2e-16 ***
#> s(year):fts_bs_s_age_bysexmale_2 12.758 15 9625.52 <2e-16 ***
#> s(year):fts_bs_s_age_bysexmale_3 14.606 15 106.51 <2e-16 ***
#> s(year):fts_bs_s_age_bysexmale_4 13.620 15 9829.84 <2e-16 ***
#> s(year):fts_bs_s_age_bysexmale_5 14.601 15 1741.23 <2e-16 ***
#> s(year):fts_bs_s_age_bysexmale_6 13.601 15 9939.09 <2e-16 ***
#> s(year):fts_bs_s_age_bysexmale_7 14.438 15 308.23 <2e-16 ***
#> s(year):fts_bs_s_age_bysexmale_8 9.738 15 12228.38 <2e-16 ***
#> s(year):fts_bs_s_age_bysexmale_9 11.218 15 72.68 <2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> R-sq.(adj) = 0.986 Deviance explained = 97.6%
#> fREML = 22150 Scale est. = 1 n = 8282Key components:
- Fixed effects:
sexcaptures baseline male-female differences,agecaptures baseline mortality rates per age - Time-varying level:
fts(year, mean_only = TRUE)models shared temporal trends
- Time-varying functions:
fts(age, by = sex)captures how age patterns evolve differently by sex
The model automatically creates a hierarchical structure where basis
functions become by variables in smooths of time, sharing
smoothing parameters for computational efficiency.
Model interpretation
Predicted functional curves
newdat <- qld_mortality
newdat$population <- 1 # Standardized predictions
newdat$preds <- predict(
mod,
newdata = newdat,
type = "response"
)
ggplot(
data = newdat,
aes(
x = age,
y = preds,
group = year,
colour = year
)
) +
geom_line() +
facet_wrap(~sex) +
scale_colour_viridis_c(name = "Year") +
labs(
x = "Age",
y = "Expected mortality rate",
title = "Model predictions: evolving mortality patterns"
) +
scale_y_log10()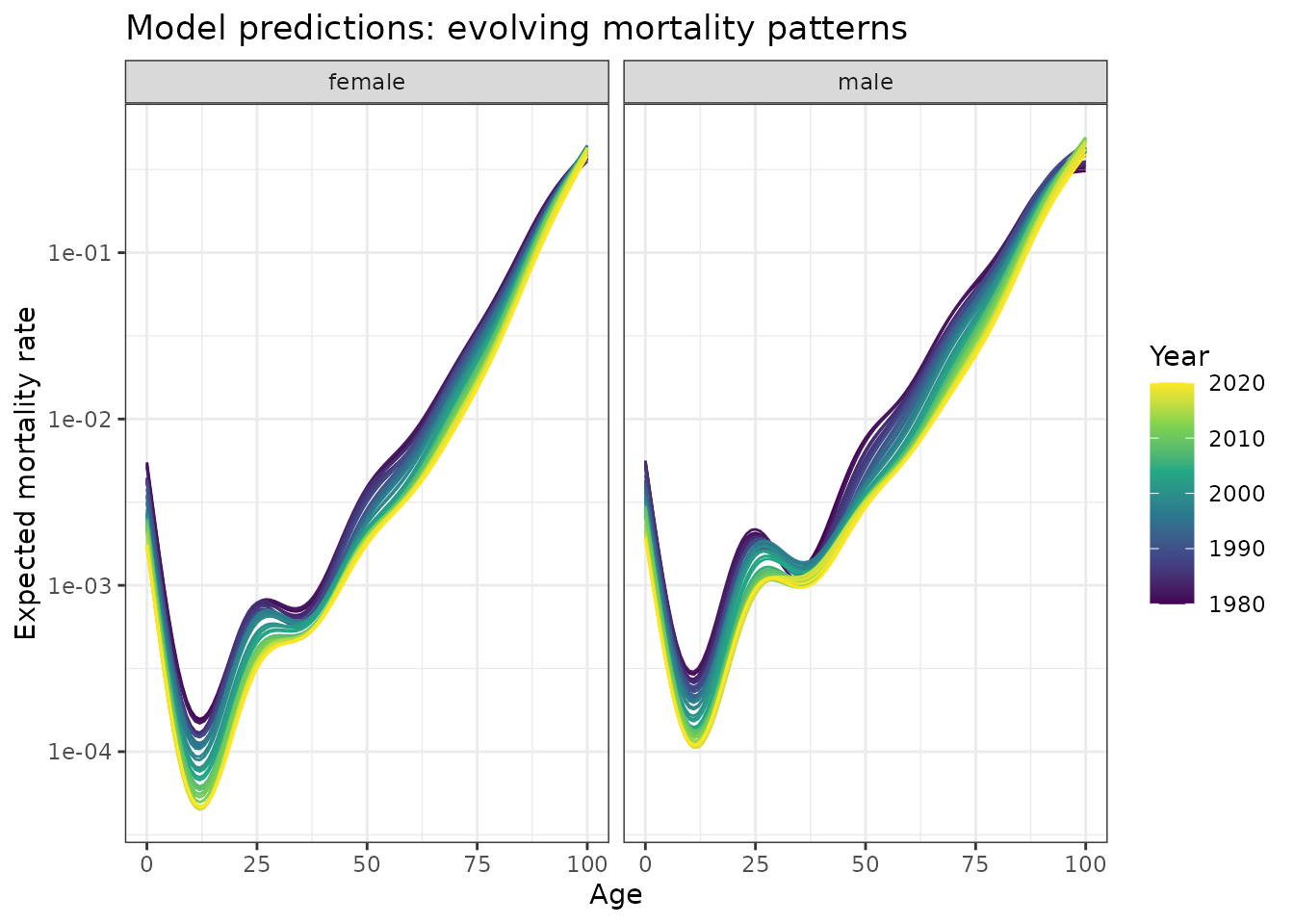
Model predictions showing expected mortality curves by age and year. Smooth curves demonstrate systematic evolution of the age-mortality relationship over four decades.
The model successfully captures the smooth evolution of J-shaped mortality curves while preserving the characteristic functional form. The downward trend across all mortality curves demonstrates substantial improvement in survival rates over the 40-year period. This reflects major advances in healthcare, improved living conditions, better nutrition, and reduced exposure to environmental hazards. Notably, the decline is not uniform across all ages - the model captures how the rate of improvement varies by age group and sex, with some periods showing more rapid mortality decline than others.
Focused analysis: teenage mortality trends
Let’s examine how mortality patterns changed for a specific age group
by leveraging the power of marginaleffects:
plot_predictions(
mod,
by = c("year", "sex"),
newdata = datagrid(
age = 17,
year = unique,
sex = unique,
population = 1
),
type = "response"
) +
labs(
x = "Year",
y = "Expected mortality rate",
title = "Teenage mortality trends over time"
) +
scale_y_log10()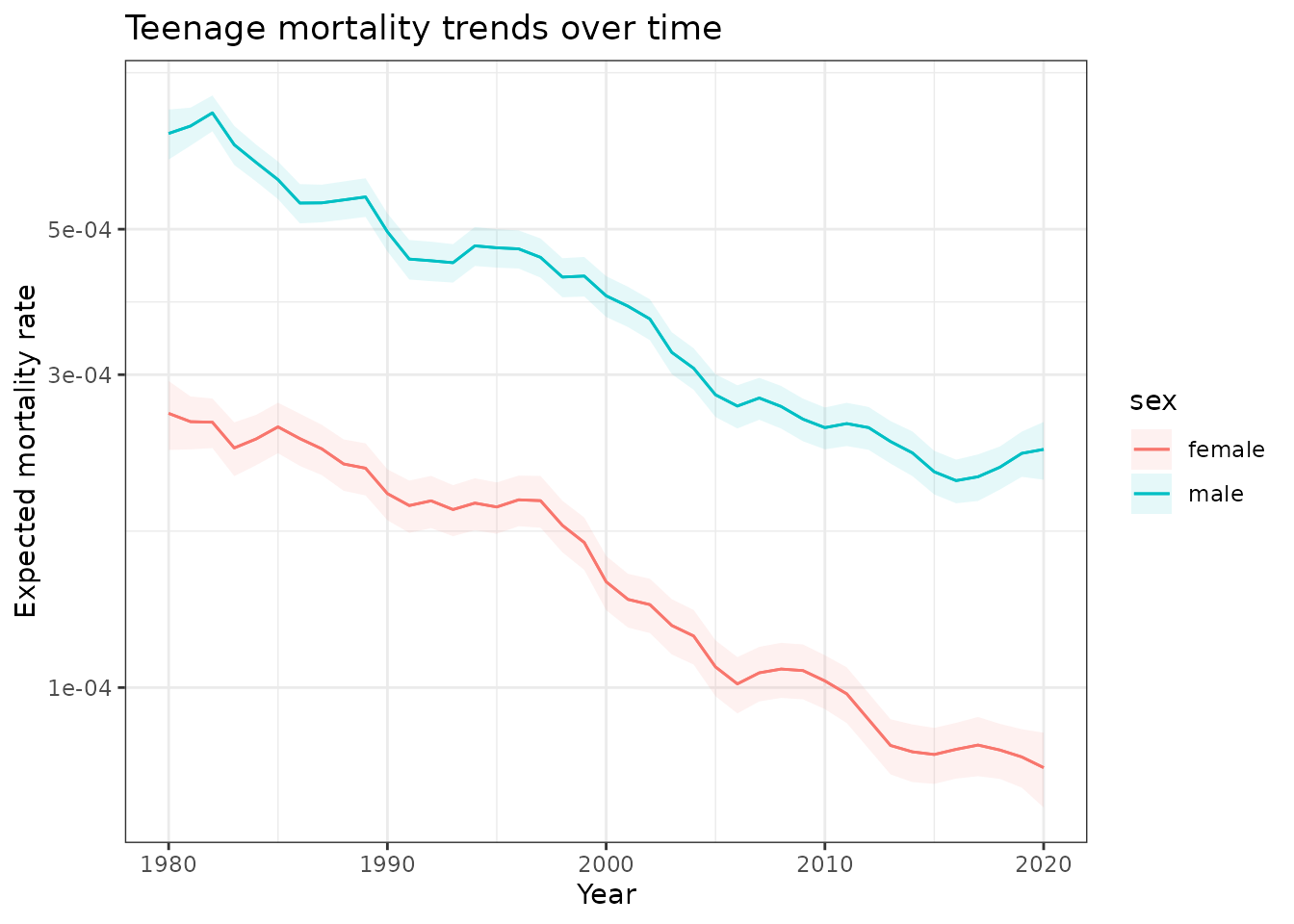
Predicted mortality trends for 17-year-olds in Queensland, 1980-2020. Shows approximately 70% decline in mortality rates with consistent male-female differences.
Rate of change analysis
We can look deeper at these predictions to understand how
fast mortality is improving, again using
marginaleffects support:
plot_slopes(
mod,
variables = "year",
by = c("year", "sex"),
newdata = datagrid(
age = 17,
year = unique,
sex = unique,
population = 1
),
type = "response"
) +
labs(
x = "Year",
y = "Rate of mortality change",
title = "Speed of mortality improvement over time"
) +
geom_hline(
yintercept = 0,
linetype = "dashed",
alpha = 0.7
)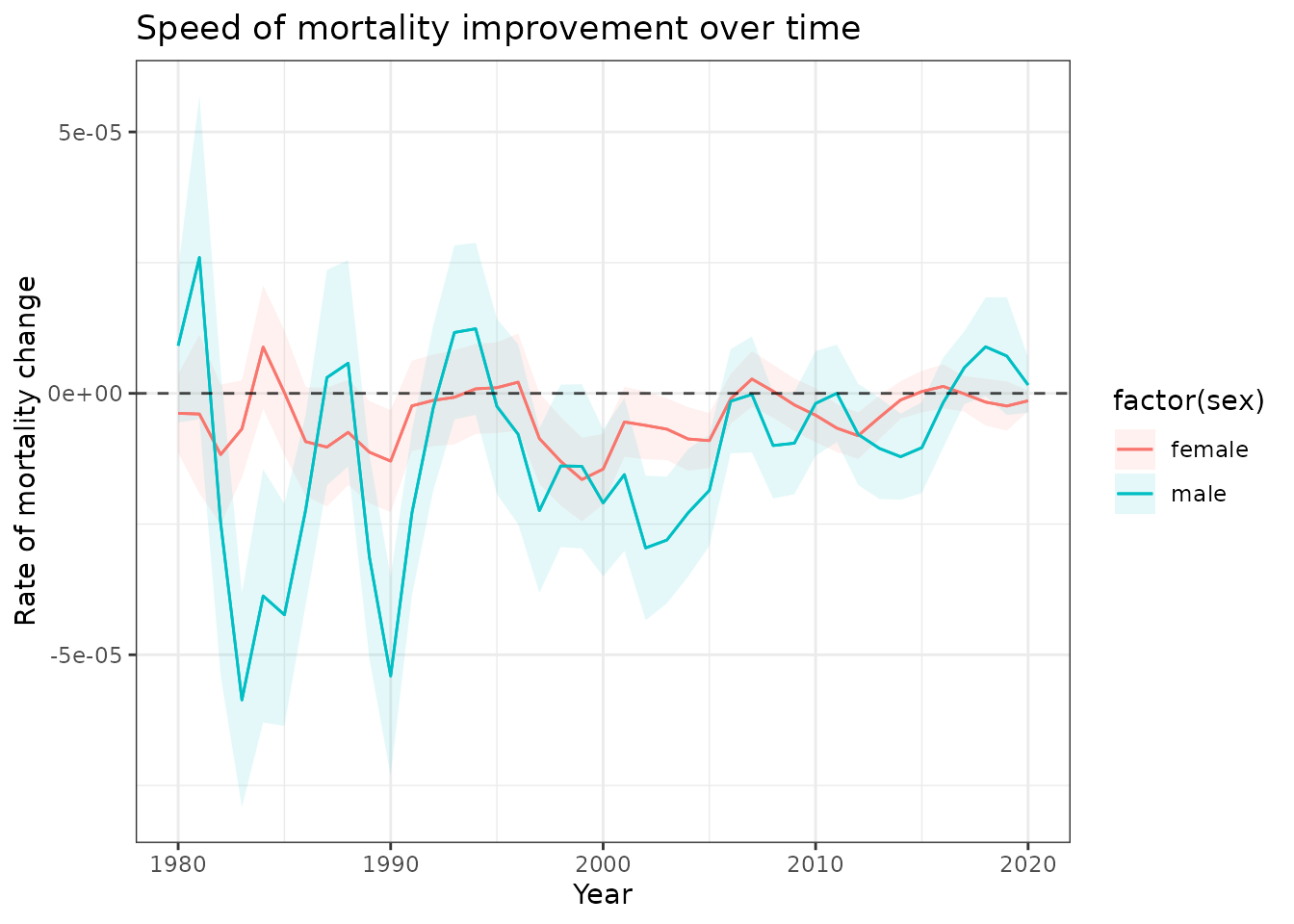
First derivatives showing the rate of mortality improvement for 17-year-olds. Fluctuations reveal periods of faster and slower mortality decline.
Extracting time-varying coefficients
The power of functional forecasting lies in treating the evolving coefficients as time series that can be forecasted:
functional_coefs <- fts_coefs(
mod,
summary = FALSE,
times = 10
)
functional_coefs
#> # A tibble: 19,475 × 6
#> .basis .parameter .time .estimate .realisation year
#> * <chr> <chr> <int> <dbl> <int> <int>
#> 1 fts_year1_mean location 1980 0.356 1 1980
#> 2 fts_year1_mean location 1981 0.368 1 1981
#> 3 fts_year1_mean location 1982 0.403 1 1982
#> 4 fts_year1_mean location 1983 0.319 1 1983
#> 5 fts_year1_mean location 1984 0.324 1 1984
#> 6 fts_year1_mean location 1985 0.318 1 1985
#> 7 fts_year1_mean location 1986 0.259 1 1986
#> 8 fts_year1_mean location 1987 0.247 1 1987
#> 9 fts_year1_mean location 1988 0.234 1 1988
#> 10 fts_year1_mean location 1989 0.265 1 1989
#> # ℹ 19,465 more rowsVisualizing coefficient evolution
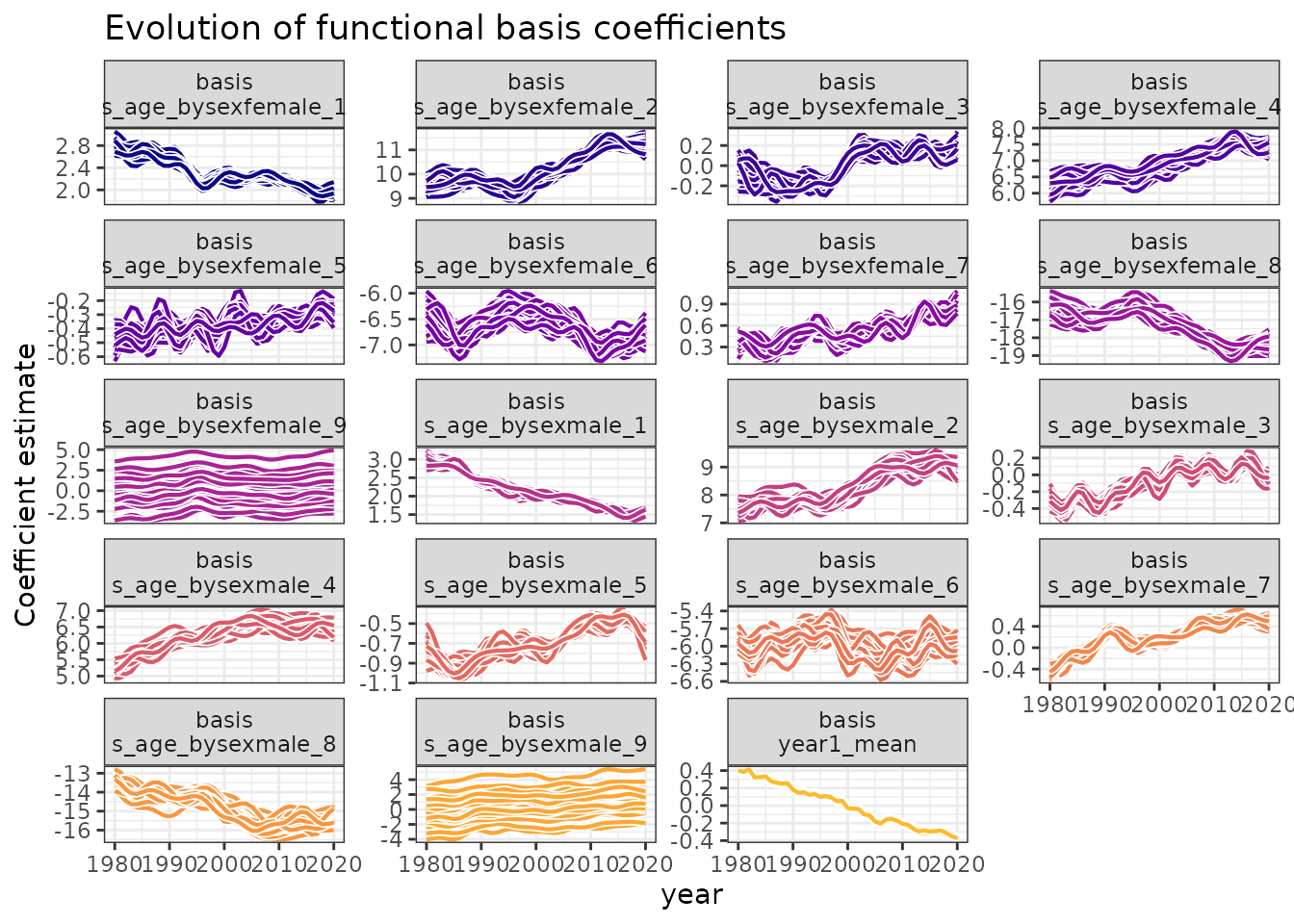
Time series of functional basis coefficients. Each panel shows how different aspects of the age-mortality relationship evolved over time, revealing complex temporal dependencies.
Interpretation:
- Complex patterns: Each coefficient series shows distinct temporal behavior
- Smooth evolution: Changes occur gradually, suitable for time series modeling
- Interdependence: Multiple coefficients work together to create the functional evolution
Forecasting mortality patterns
Coefficient forecasting
functional_fc <- forecast(
object = functional_coefs,
h = 5, # 5-year horizon
times = 10, # Forecast replicates
model = "ARIMA"
)
functional_fc
#> # A tsibble: 59,375 x 6 [1Y]
#> # Key: .basis, .realisation, .model, .rep [11,875]
#> .basis .realisation .model year .rep .sim
#> <chr> <int> <chr> <dbl> <chr> <dbl>
#> 1 fts_bs_s_age_bysexfemale_1 1 ARIMA 2021 1 1.96
#> 2 fts_bs_s_age_bysexfemale_1 1 ARIMA 2022 1 1.92
#> 3 fts_bs_s_age_bysexfemale_1 1 ARIMA 2023 1 1.86
#> 4 fts_bs_s_age_bysexfemale_1 1 ARIMA 2024 1 1.81
#> 5 fts_bs_s_age_bysexfemale_1 1 ARIMA 2025 1 1.78
#> 6 fts_bs_s_age_bysexfemale_1 1 ARIMA 2021 10 1.96
#> 7 fts_bs_s_age_bysexfemale_1 1 ARIMA 2022 10 1.93
#> 8 fts_bs_s_age_bysexfemale_1 1 ARIMA 2023 10 1.88
#> 9 fts_bs_s_age_bysexfemale_1 1 ARIMA 2024 10 1.86
#> 10 fts_bs_s_age_bysexfemale_1 1 ARIMA 2025 10 1.86
#> # ℹ 59,365 more rowsThe forecast includes uncertainty in both the time series models and the original coefficient estimation, providing realistic prediction intervals.
Future mortality patterns
Let’s demonstrate how to generate complete mortality curve forecasts by combining the forecasted coefficients with the model structure. Here we will use the ensemble model (“ENS”) to forecast the time-varying mean, which tends to give more robust forecasts than any of the mdoels on their own:
# Create forecast data for future years
future_years <- 2021:2025
newdata_forecast <- expand.grid(
age = unique(qld_mortality$age),
sex = unique(qld_mortality$sex),
year = future_years,
population = 1
)
# Generate forecasted mortality curves using forecast()
# This integrates the time series forecasts of coefficients
mortality_forecasts <- forecast(
object = mod,
newdata = newdata_forecast,
model = "ARIMA",
mean_model = "ENS",
type = "expected"
)
head(mortality_forecasts)
#> # A tibble: 6 × 6
#> .estimate .error .q2.5 .q10 .q90 .q97.5
#> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 0.00167 0.00161 0.00107 0.00130 0.00236 0.00318
#> 2 0.00105 0.00109 0.000683 0.000776 0.00125 0.00204
#> 3 0.000665 0.00142 0.000431 0.000512 0.000788 0.00104
#> 4 0.000446 0.00226 0.000285 0.000358 0.000558 0.000747
#> 5 0.000285 0.00232 0.000190 0.000216 0.000360 0.000428
#> 6 0.000194 0.00141 0.000113 0.000144 0.000241 0.000330
# Plot forecasted mortality rates, together with uncertainties
ggplot(mortality_forecasts |>
dplyr::bind_cols(newdata_forecast),
aes(x = age,
y = .estimate,
group = year,
colour = year)) +
geom_ribbon(aes(
ymin = .q10,
ymax = .q90,
fill = year
),
alpha = 0.2,
colour = NA) +
geom_line() +
facet_wrap(~sex) +
scale_colour_viridis_c(name = "Year") +
scale_fill_viridis_c(name = "Year") +
labs(
x = "Age",
y = "Mortality rate",
title = "Expected mortality"
) +
scale_y_log10()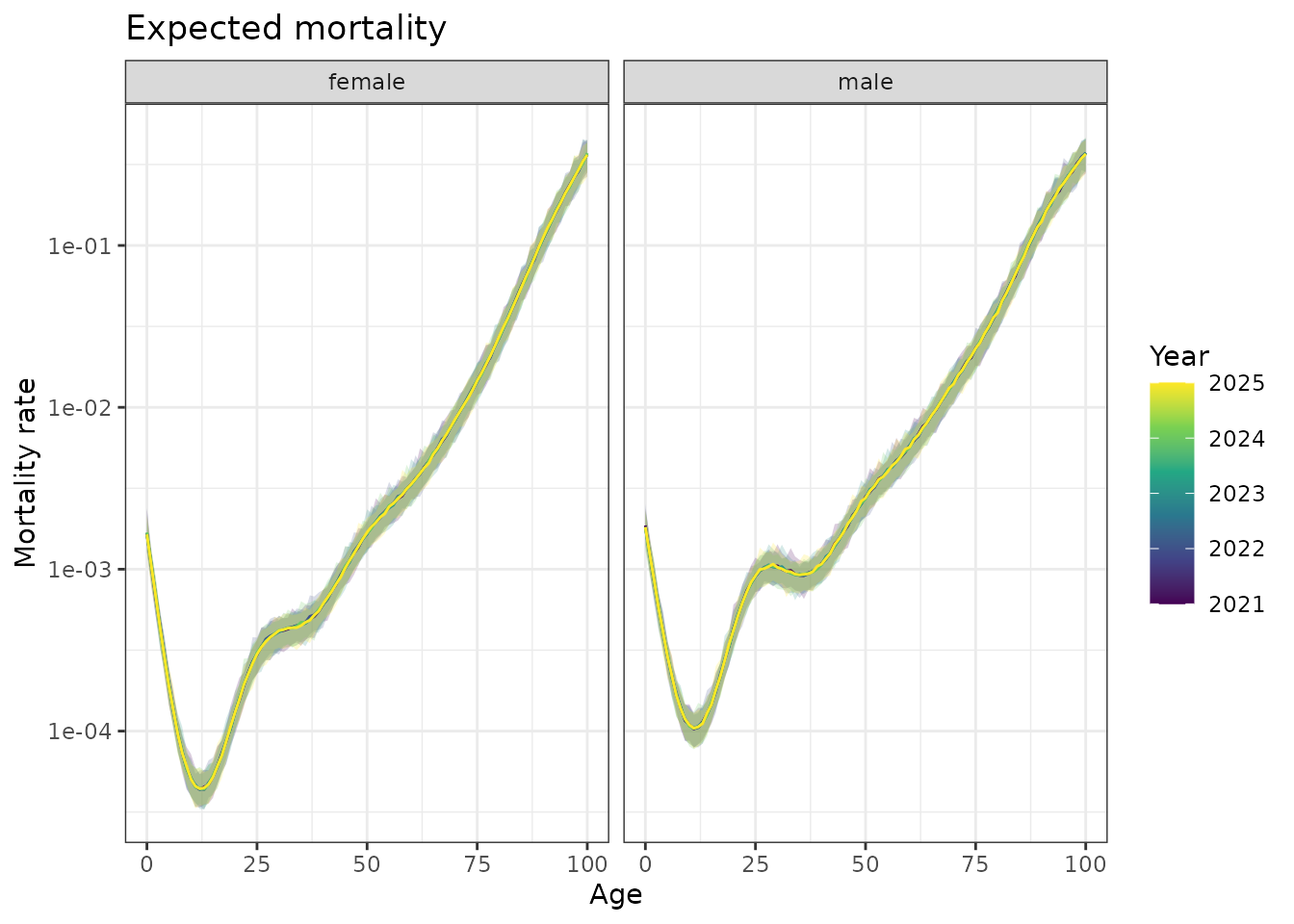
Forecasted mortality curves for Queensland, 2021-2025. Shows how the functional forecasting approach predicts future age-mortality patterns.
This forecasting approach enables prediction of entire functional
relationships by combining time series forecasts of the functional
coefficients with the underlying model structure. We could easily then
convert this to a fable object (using
as_fable()) and compare with holdout data to compute any of
the forecast scores supported by fable.
Key insights and methodology
What we learned about Queensland mortality
- Systematic improvement: Mortality declined across all age groups over 40 years
- Functional evolution: Changes involved curve shape, not just magnitude shifts
- Heterogeneous trends: Rate of improvement varied by age and sex
- Complex patterns: Simple parametric models would miss these relationships
Why functional forecasting matters
- Captures complexity: Models evolving functional relationships traditional methods miss
- Preserves structure: Maintains biological/physical constraints in
forecasts
- Quantifies uncertainty: Propagates uncertainty through the entire functional evolution
- Policy relevance: Enables sophisticated demographic projections
Model design principles
- Hierarchical structure: Shared trends with group-specific deviations
- Temporal smoothing: Balance flexibility with smoothness using
time_kandtime_m - Basis choice: Thin plate splines work well for age and time
- Computational efficiency:
bam()engine handles large datasets effectively
Conclusion
Functional forecasting with ffc provides a principled
approach to modeling and predicting evolving functional relationships.
By treating time-varying coefficients as forecastable time series, we
can:
- Capture complexity that traditional methods miss
- Generate realistic forecasts of entire functional
relationships
- Quantify uncertainty appropriately across all sources
- Provide actionable insights for policy and planning
The Queensland mortality analysis demonstrates these capabilities in a real-world context, revealing patterns and trends that inform our understanding of demographic change and enable sophisticated projection methods for public health planning.